The project aims to utilize realistic bacterial membrane models for experimental studies on permeation and interaction with antimicrobial agents at atomic resolution.
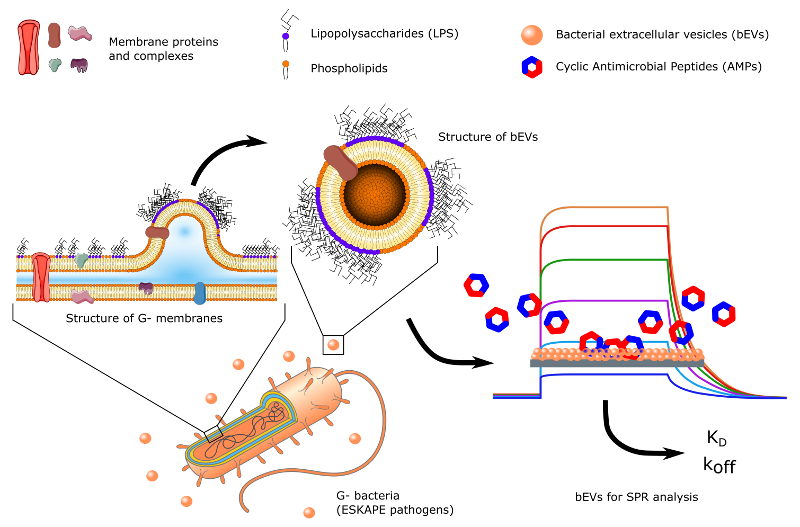
Antibiotic resistance is a dangerously rising problem of the modern world. According to the WHO this serious threat is no longer a prediction for the future, it is happening right now in every region of the world and has the potential to affect anyone, of any age, in any country. Antibiotic resistance — is now a major threat to public health. Misuse and overuse of antimicrobials together with somehow insufficient infection control and prevention of infections led to accelerated development of different mechanisms of bacterial resistance to the antibiotics. To cope with this problem, it is essential to keep searching for the new antimicrobial agents and design new antibiotics. This in turn is sorely dependent on scrutinization of the mechanisms of the acquired antimicrobial resistance. Thus, the project aims to develop a novel in vitro nanoparticle based system for direct experimental studies of truly realistic bacterial membrane models for permeation and interaction with antimicrobial agents at atomic resolution. To achieve the aim the project utilizes ability of bacteria to produce extracellular vesicles (referred to as bacterial ectracellular vesicles (bEV)). Provoking isolates of resistant and non-resistant strains of Gram negative (G-) and Gram positive (G+) bacteria to export sufficient amounts of MV will allow assembly of the close to native barrier permeation and interaction models uitlizing analytical techniques as SPR and NMR.
[Loading...]